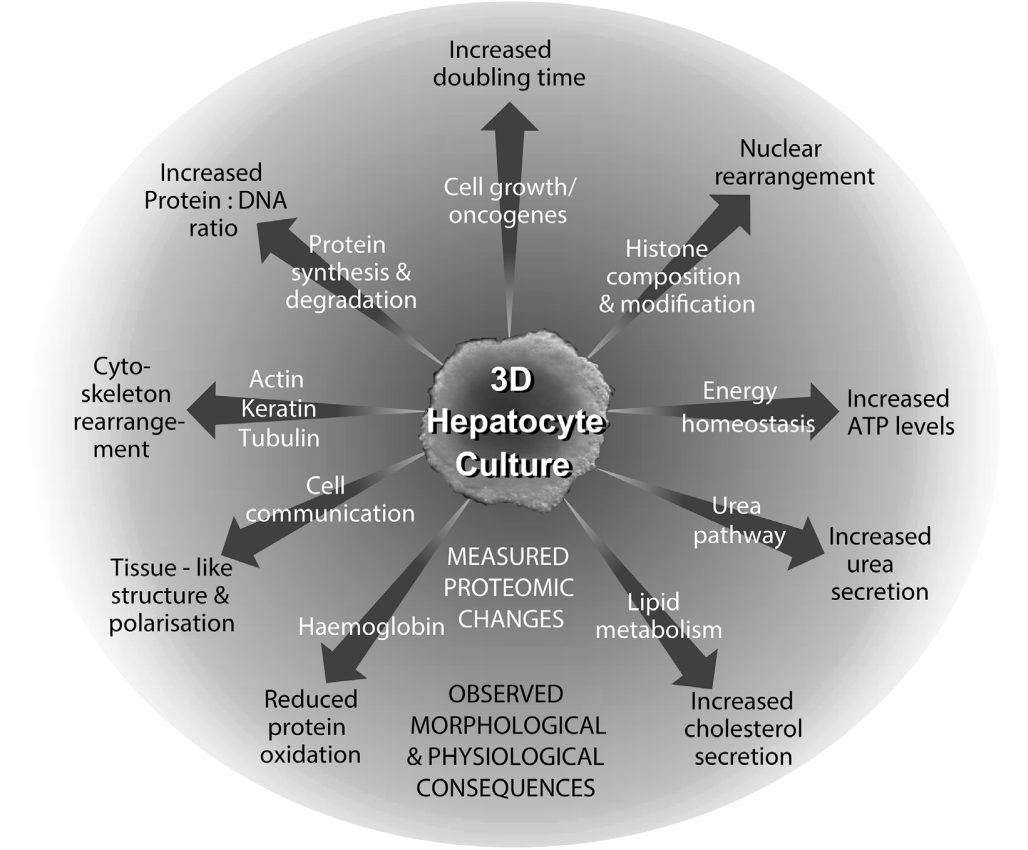
Celvivo culture characteristics
The CelVivo system creates an environment which promotes the growth and maintenance of large 3D tissue mimetic structures, whether they are spheroids, organoids or other aggregates.
There are several advantages of using a clinostat for cell culture. The two main benefits of the clinostat are low shear stress and active diffusion. These benefits result in the cells being able to present a physiological performance similar to that seen in vivo and the reestablishment of intercellular communication. Some of the physiological changes observed in the cell culture when cultivated with the CelVivo Technology are described below. These and additional data are presented in the cited peer-reviewed publications.
Characteristics of cells cultivated in ClinoStar
These observations are made using the C3A derivative of the HepG2 hepatocytes (ATCC CRL-10741) cultivated for at least 18 days in the clinostat culture. These cells need 18 days to recover from trypsin-induced damage and exhibit an in vivo physiological performance. This is not special for this cell line and there are numerous publications describing that other cell lines need similar recovery (or ‘maturation’ periods). Once recovered the cells remain stable and can be used for experimetns for at least 24 days, and often longer. In one experiment a spheroid was maintained in the system 302 days (Wrzesinski et al. 2013 )
Proliferation
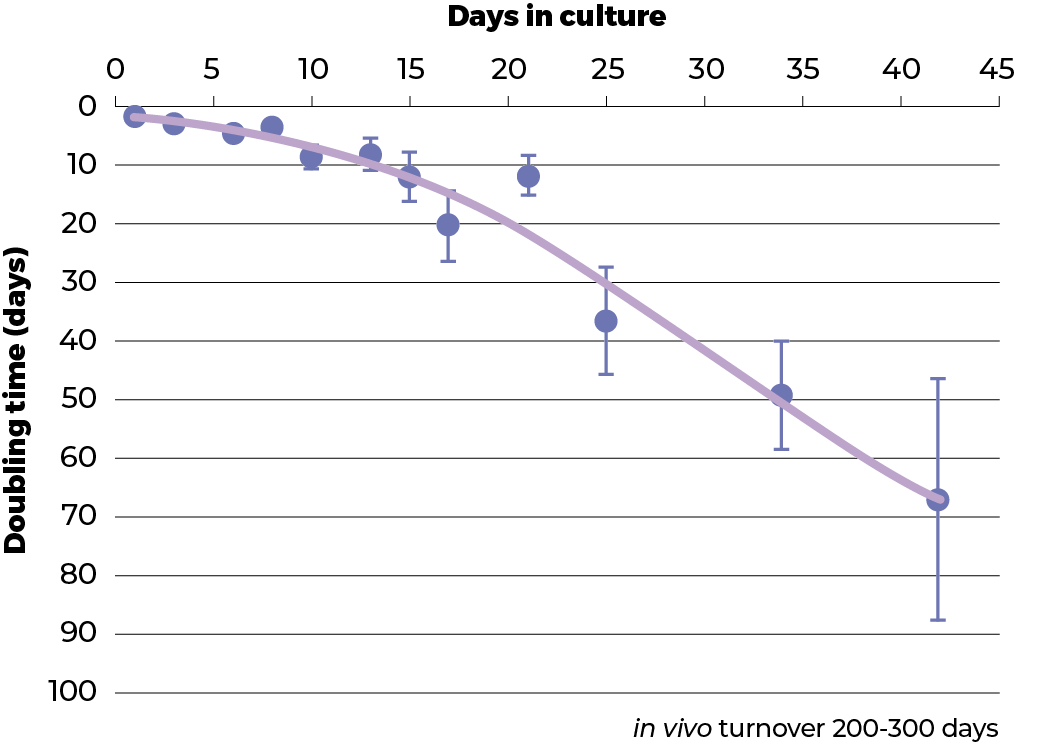
The growth rate of the spheroids was evaluated using the protein content as an indicator of the number of cells present,. In classical monolayer cell culture conditions (‘2D’) the C3A cell line has a doubling time of about 1 day. This is much faster than even the tumour from which it was derived. In clinostat culture the proliferation of the C3A cells was shown to dramatically decrease during a 42 day culture period so that after the 42nd day, the doubling time was about 65 days. This is approaching the 200-300 day turnover rate of hepatocytes in the healthy liver.
Spheroid viability
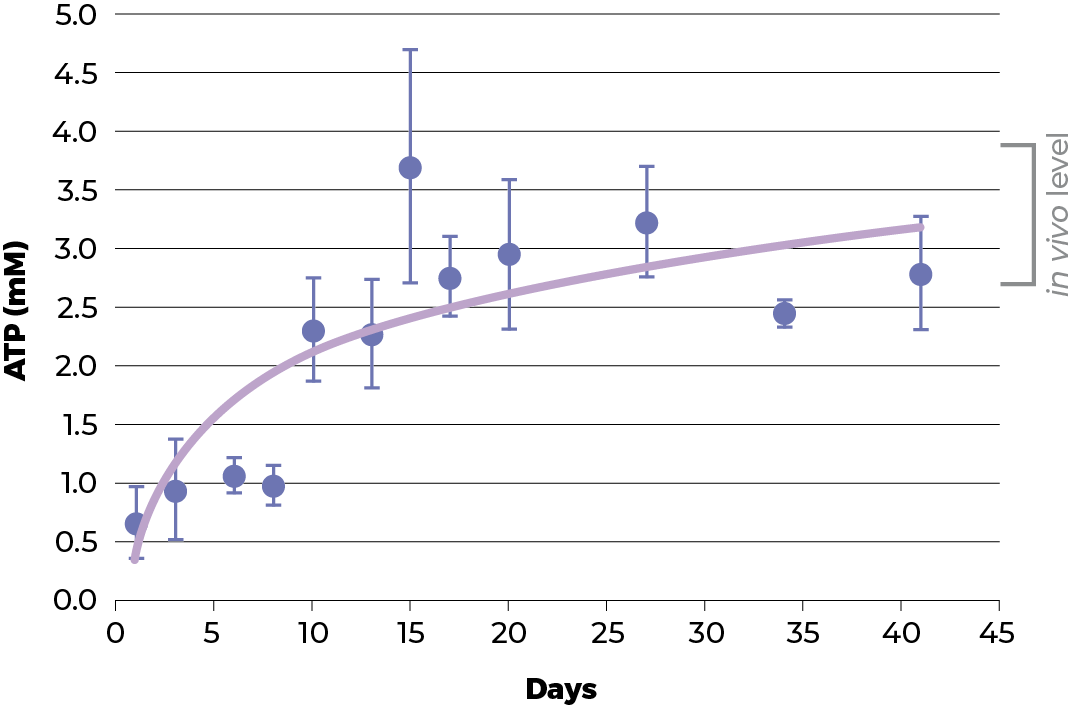
As the proliferation rate decreases, one could argue that the cells have died or decreased their metabolic activity. ATP levels are a good indicator of cell viability and metabolic activity. In HepG2 spheroids the ATP content increases as the spheroids in the culture recover. This illustrates that the cells are not dying or have not reduced their metabolic activity but are in fact probably more healthy than before. This observation is substantiated by an essentially unchanged rate of cell death (as estimate by adenylate kinase release).
Physiological performance
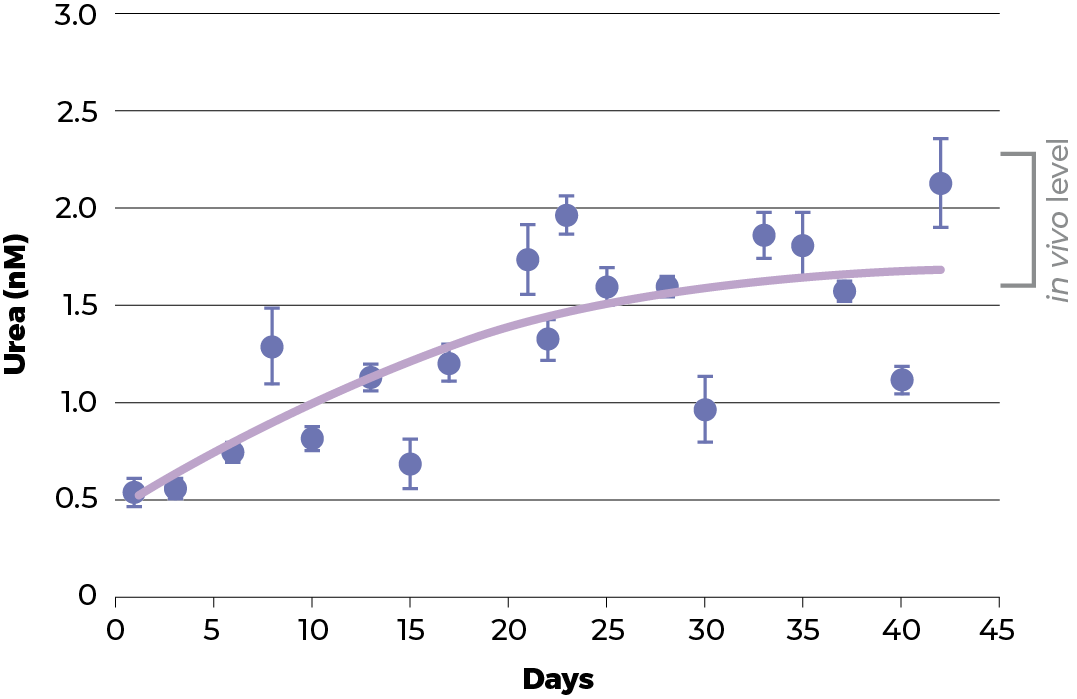
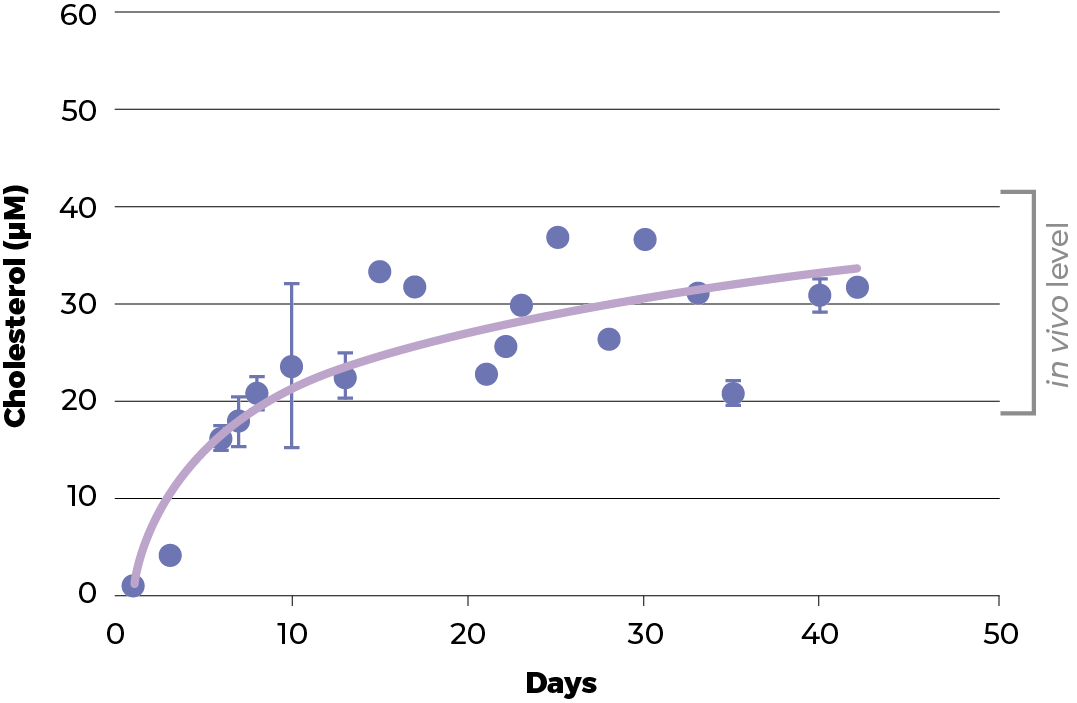
To have a good model, the physiology functionality and structure of the spheroids should resemble its parental tissue. Therefore, the functionality of C3A spheroids culture was evaluated and benchmarked against two normal physiological functions of healthy liver hepatocytes: urea and cholesterol production.
During the recovery period of the spheroids the level of urea and cholesterol increase to those seen in vivo. Around day 18, the level of both urea and cholesterol start to plateau and thereafter remain stable until at least day 42 (when the experiment was terminated).
Ultrastructure
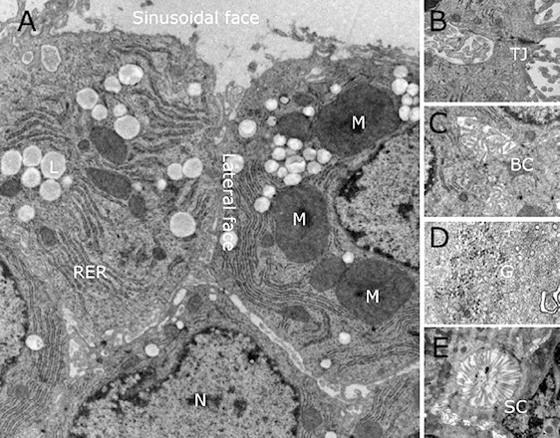
Examined by electron microscopy, cells in 3D spheroids exhibit all the ultrastructural organelles normally seen in eukaryotic cells (nuclei, mitochondria, rough endoplasmic reticulum, etc. (N, M, RER respectively). Cells exhibit clear tight junctions (TJ) that compartmentalize the plasma membrane, allowing different receptors, transporters and enzyme systems to be sequestered to specific regions. In addition, spheroids exhibit several features that are infrequently seen when the C3A cells are grown under traditional monolayer cell culture conditions (2D). These features include the formation of bile canaliculi-like structures in the microvilli-free inter-hepatocyte faces. In vivo, in the liver these canaliculi merge and develop into bile ducts leading to the gall bladder. Microvilli can be seen in the sinusoidal-like channels. In vivo, these channels, together with an incomplete layer of endothelial cells, form the space of Disse. Spheroids also exhibit glycogen granules characteristic for the storage of glucose in the liver
Epigenetics
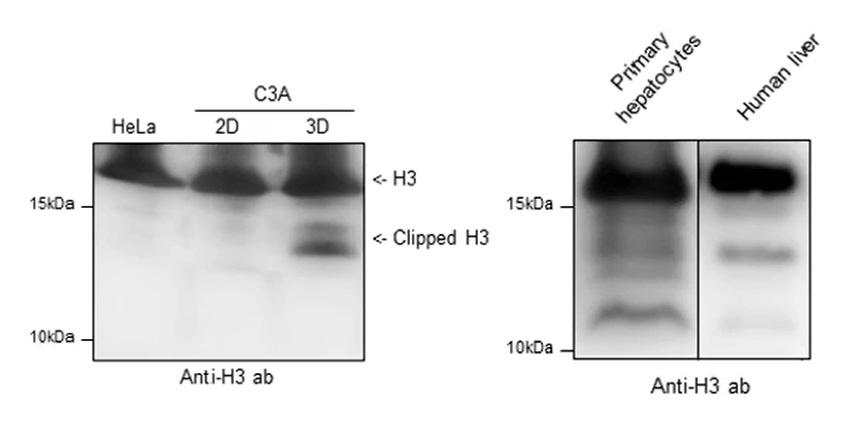
Post-translational modifications (PTMs) of histone proteins play a fundamental role in regulation of DNA-templated processes. There is also growing evidence that proteolytic cleavage of histone N-terminal tails, known as histone clipping, influences nucleosome dynamics and functional properties. Using top-down and middle-down protein analysis by mass spectrometry, C3A spheroids grown in clinostat 3D culture recover histone H2B and H3 N-terminal tail clipping and illustrate a relationship between clipping and co-existing PTMs of histone H3. Histones H2B and H3 undergo proteolytic processing in primary human hepatocytes and the hepatocellular carcinoma cell line HepG2/C3A when grown in spheroid (3D) culture, but not when the same cell line is grown in a flat (2D) culture.
Metabolic reprogramming
Compared to cells cultivated using the classical (‘2D’) cell culture techniques, cells cultivated as spheroids in ClinoStar for 21 days show wide-ranging metabolic and architectural alterations necessary to recover the in vivo physiological performance.
A proteomic study of C3A cells grown as traditional monolayer (2D) or 3 dimensional clinostat spheroid culture (3D) showed a significant metabolic reprogramming resulting in structurally changes in actin organization, an increase in microtubules and a decrease in keratins 8 and 18. Metabolically, glycolysis, fatty acid metabolism and the pentose phosphate shunt were increased while the TCA cycle and oxidative phosphorylation was unchanged. Enzymes involved in cholesterol and urea synthesis were increased consistent with the attainment of cholesterol and urea production rates seen in vivo. DNA repair enzymes were increased even though cells were predominantly in G1/Go. Transport around the cell – along the microtubules, through the nuclear pore and in various types of vesicles was prioritized. There were numerous coherent changes in transcription, splicing, translation, protein folding and degradation. Even the amounts of individual proteins within complexes were shown to be changed in a highly coordinated manner.
In conclusion, the biochemistry occurring within cells in 3D culture is very different to that occurring in 2D cultures. The take home message is that one cannot use data produced in 2D cultures to understand the in vivo situation. 3D cultures mimic the in vivo situation and produce useful, valuable data